BIOT
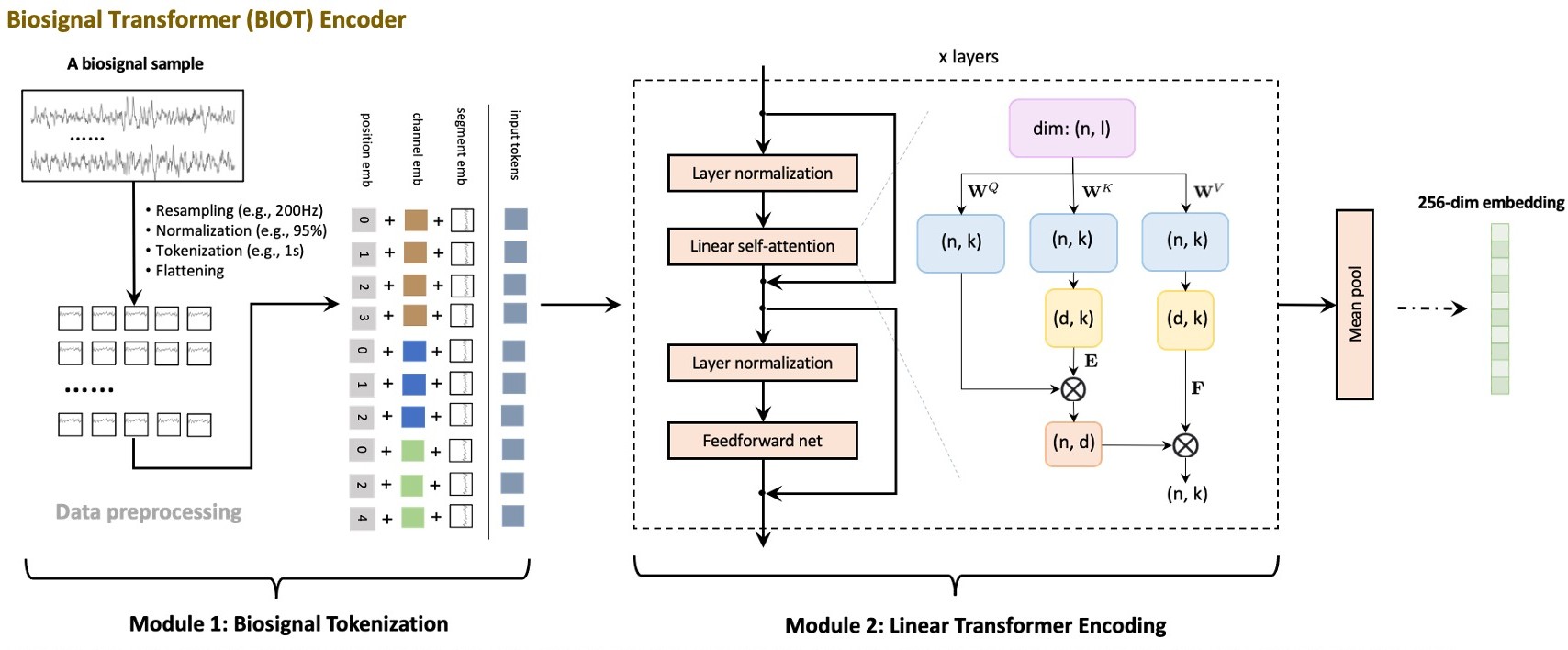
BIOT from Yang et al (2023) [Yang2023]
Architecture-only repository. Documents the
braindecode.models.BIOTclass. No pretrained weights are distributed here. Instantiate the model and train it on your own data.
Quick start
pip install braindecode
from braindecode.models import BIOT
model = BIOT(
n_chans=16,
sfreq=200,
input_window_seconds=10.0,
n_outputs=2,
)
The signal-shape arguments above are illustrative defaults — adjust to match your recording.
Documentation
- Full API reference: https://braindecode.org/stable/generated/braindecode.models.BIOT.html
- Interactive browser (live instantiation, parameter counts): https://huggingface.co/spaces/braindecode/model-explorer
- Source on GitHub: https://github.com/braindecode/braindecode/blob/master/braindecode/models/biot.py#L56
Architecture
Parameters
| Parameter | Type | Description |
|---|---|---|
embed_dim |
int, optional | The size of the embedding layer, by default 256 |
num_heads |
int, optional | The number of attention heads, by default 8 |
num_layers |
int, optional | The number of transformer layers, by default 4 |
activation: nn.Module, default=nn.ELU |
— | Activation function class to apply. Should be a PyTorch activation module class like nn.ReLU or nn.ELU. Default is nn.ELU. |
return_feature: bool, optional |
— | Changing the output for the neural network. Default is single tensor when return_feature is True, return embedding space too. Default is False. |
hop_length: int, optional |
— | The hop length for the torch.stft transformation in the encoder. The default is 100. |
sfreq: int, optional |
— | The sfreq parameter for the encoder. The default is 200 |
References
- Yang, C., Westover, M.B. and Sun, J., 2023, November. BIOT: Biosignal Transformer for Cross-data Learning in the Wild. In Thirty-seventh Conference on Neural Information Processing Systems, NeurIPS.
- Yang, C., Westover, M.B. and Sun, J., 2023. BIOT Biosignal Transformer for Cross-data Learning in the Wild. GitHub https://github.com/ycq091044/BIOT (accessed 2024-02-13)
Citation
Cite the original architecture paper (see References above) and braindecode:
@article{aristimunha2025braindecode,
title = {Braindecode: a deep learning library for raw electrophysiological data},
author = {Aristimunha, Bruno and others},
journal = {Zenodo},
year = {2025},
doi = {10.5281/zenodo.17699192},
}
License
BSD-3-Clause for the model code (matching braindecode). Pretraining-derived weights, if you fine-tune from a checkpoint, inherit the licence of that checkpoint and its training corpus.